 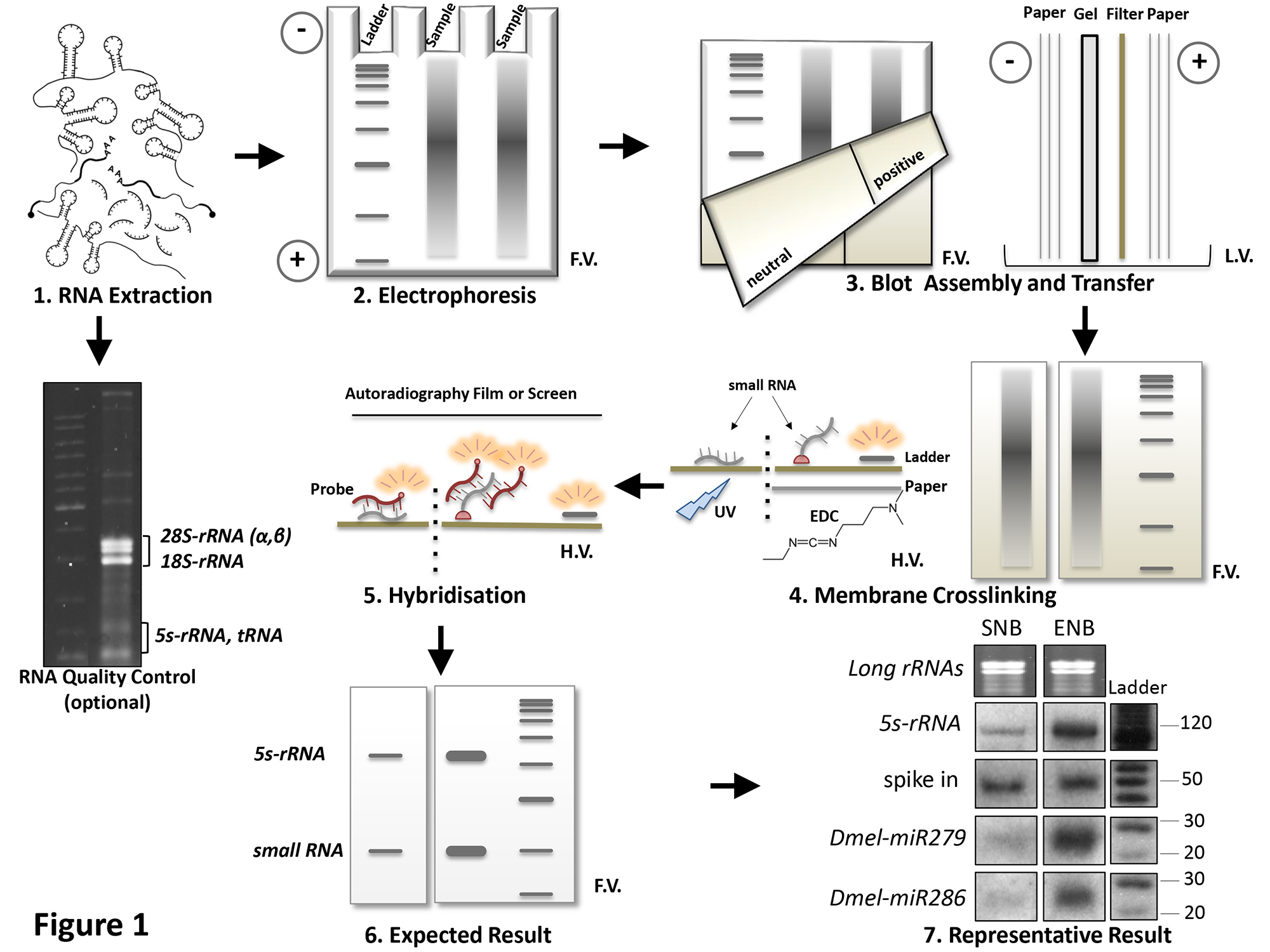
Thus the reverse procedure, though originally uncommon, enabled the one-at-a-time study of gene expression using northern analysis to evolve into gene expression profiling, in which many (possibly all) of the genes in an organism may have their expression monitored. The following steps are described in detail: (1) preparation of highly labelled biotinylated-probes generated by polymerase chain reaction (PCR), (2) sample preparation for gel electrophoresis, (3) blotting of the electrophoretically. The use of DNA microarrays that have come into widespread use in the late 1990s and early 2000s is more akin to the reverse procedure, in that they involve the use of isolated DNA fragments affixed to a substrate, and hybridization with a probe made from cellular RNA. In this chapter we describe one nonradioactive method for Northern blot analysis using biotinylated probes. In this procedure, the substrate nucleic acid (that is affixed to the membrane) is a collection of isolated DNA fragments, and the probe is RNA extracted from a tissue and radioactively labelled. The Northern blot is similar to the Southern blot except that RNA instead of DNA is the subject of analysis in this technique. For smaller fragments denaturing polyacrylamide urea gels are employed.Īs in the Southern blot, the hybridization probe may be made from DNA or RNA.Ī variant of the procedure known as the reverse northern blot was occasionally (although, infrequently) used. A notable difference in the procedure in case of agarose gels, (as compared with the Southern blot) is the addition of formaldehyde which acts as a denaturant. 2 in a Stratalinker UV crosslinker (Stratagene), the membrane was blocked with 2 Block-Ace (Dainihon Pharmaceutical) in TBS-T (50 mM Tris, 150 mM sodium chloride, and 0.1 Tween 20, pH 7.4) for 1 h at room temperature and then incubated with the primary antibodies (diluted 1:500 for anti-m 1 A, anti-PU and anti-m. The gels may be run on either agarose or denaturing polyacrylamide gels depending on the size of the RNA to be detected. Northern blot analysis can be used to investigate whether a gene is overexpressed or underexpressed in cancer cells as compared to normal cells. This technique was developed in 1977 by James Alwine, David Kemp, and George Stark at Stanford University. 2 This technique was named Northern blottingas a pun on Southern’s name. It takes its name from the similarity of the procedure to the Southern blot procedure, named for biologist Edwin Southern, used to study DNA, with the key difference that, in the northern blot, RNA, rather than DNA, is the substance being analyzed by electrophoresis and detection with a hybridization probe. In 1977, gene expression analysis ascended dramatically when George Stark and colleagues replicated the con guration of Southern’s transfer apparatus in an effort to transfer cellular RNA to chemically activated cellulose paper. The northern blot is a technique used in molecular biology research to study gene expression.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |



 RSS Feed
RSS Feed
